Current Research Areas & Applications
Material Systems of Interest
Our biofunctional materials are based on sol-gel condensation and chemically and physically induced polymerization of protein-doped and peptide-modified polymer composites, which enable engineered materials and devices to communicate biochemically with living cells. We have developed strong expertise in integration of proteins and cells into fabricated materials including inorganic oxides, such as silica, and organic polymers, such as collagen and polyethers. We further enhanced the biofunctionality of these materials by nanostructuring the metal interfaces (e.g. nanostructured platinum and carbon nanotubes) to improve the sensitivity of biomolecule-based biosensors and the biointerface surface (e.g. nanorough silica) to improve the biocompatibility of the material-cell interface.
Current Projects
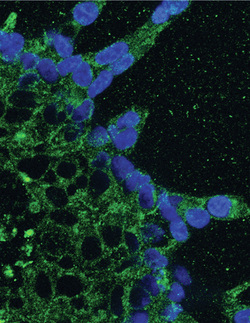
Glioblastoma - Grade IV Brain Tumors:Tissue Engineered 3D Brain Mimics for Glioblastoma Research and Therapy Development
Glioblastoma is a deadly primary brain cancer that remains highly resistant to current treatment of combined surgery, chemotherapy and radiation. In patients the cancer cells migrate to invade the entire brain making surgical removal of tumors only a temporary solution. In addition, the cancer cells resist chemotherapy and radiation. Drug testing conducted with traditional culture methods on 2D dishes has proven a poor predictor of drug efficacy in human patients; the organ context and brain tumor microenvironment may play an important role in how the cancer cells invade healthy brain regions and leverage protective strategies to resist treatment. In order to understand the interaction between the brain tissue and the cancer cells and to develop new strategies to outsmart the cancer cells, we are developing a multi-cell type and multi-niche tissue engineered brain construct for 3D culture of glioblastoma. The brain constructs consist of multiple extracellular matrix components including oligomeric collagen and hyaluronan to match the varied mechanical and chemical microenvironments in the brain. We also engineer pseuodo blood vessels and include normal glial and endothelial cells, which can alter and enhance cell migration response to chemotherapy.
Funded by: IUPUI Center for the Cure of Glioblastoma
Recent Publications: Herrera-Perrez et al. 2015 Tissue Engineering Part A
Glioblastoma is a deadly primary brain cancer that remains highly resistant to current treatment of combined surgery, chemotherapy and radiation. In patients the cancer cells migrate to invade the entire brain making surgical removal of tumors only a temporary solution. In addition, the cancer cells resist chemotherapy and radiation. Drug testing conducted with traditional culture methods on 2D dishes has proven a poor predictor of drug efficacy in human patients; the organ context and brain tumor microenvironment may play an important role in how the cancer cells invade healthy brain regions and leverage protective strategies to resist treatment. In order to understand the interaction between the brain tissue and the cancer cells and to develop new strategies to outsmart the cancer cells, we are developing a multi-cell type and multi-niche tissue engineered brain construct for 3D culture of glioblastoma. The brain constructs consist of multiple extracellular matrix components including oligomeric collagen and hyaluronan to match the varied mechanical and chemical microenvironments in the brain. We also engineer pseuodo blood vessels and include normal glial and endothelial cells, which can alter and enhance cell migration response to chemotherapy.
Funded by: IUPUI Center for the Cure of Glioblastoma
Recent Publications: Herrera-Perrez et al. 2015 Tissue Engineering Part A
High Density Array Biosensors for Spatial Mapping of Cellular Gradients and Flux
The flux and resulting concentration gradients of small molecules and ions at the surface of cells carry important information that is communicated both internally to the cell and externally to neighboring cells. Examples of such important cellular flux in biology include neurotransmitter release and reuptake in neurons, glucose transport for adapted cell metabolism in cancer cells, and ion flux in plant gravity sensing during development. Self-referencing biosensors are currently the best method for quantifying cellular flux dynamics, but this method is limited to measurements at one location in space on one cell at a time. In addition, this method is not portable and cannot be used for sensing in remote locations such as on space missions, in environmental monitoring, or in high throughput biomedical applications. No method currently exists that can quantify the spatial map and dynamics of true cellular flux and biomolecule gradients from single cells in a chip format.
This research integrates fundamental advances in electrode array design, signal detection, and biorecognition molecule confinement at the nano and microscales to create a platform for basic biological experiments in extracellular signaling. The overall objective of this work is to create a technology for the spatial and temporal mapping of biomolecule gradients and flux at the level of single cells by creating 1D and 2D arrays of individually addressable nano/microscale electrochemical sensors.
Funded by: National Science Foundation NanoBiosensing Program
The flux and resulting concentration gradients of small molecules and ions at the surface of cells carry important information that is communicated both internally to the cell and externally to neighboring cells. Examples of such important cellular flux in biology include neurotransmitter release and reuptake in neurons, glucose transport for adapted cell metabolism in cancer cells, and ion flux in plant gravity sensing during development. Self-referencing biosensors are currently the best method for quantifying cellular flux dynamics, but this method is limited to measurements at one location in space on one cell at a time. In addition, this method is not portable and cannot be used for sensing in remote locations such as on space missions, in environmental monitoring, or in high throughput biomedical applications. No method currently exists that can quantify the spatial map and dynamics of true cellular flux and biomolecule gradients from single cells in a chip format.
This research integrates fundamental advances in electrode array design, signal detection, and biorecognition molecule confinement at the nano and microscales to create a platform for basic biological experiments in extracellular signaling. The overall objective of this work is to create a technology for the spatial and temporal mapping of biomolecule gradients and flux at the level of single cells by creating 1D and 2D arrays of individually addressable nano/microscale electrochemical sensors.
Funded by: National Science Foundation NanoBiosensing Program
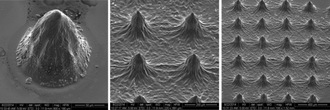

Multi-Optrode Arrays (MOAs) for Quantification of Biological Gradients
We are using light printing methods to create 2 and 3D arrays of biologically active optrodes using composites of biological glasses, photo-crosslinkable polymers and genetically encoded molecular sensors. These sensor arrays are designed to measure gradients of extracellular signals (e.g. neurotransmitters, calcium, inflammatory cytokines) and metabolites (e.g. oxygen, glucose, lactate) from tumor spheroids, developing embryos, pancreatic islets and networks of neural cells.
We are using light printing methods to create 2 and 3D arrays of biologically active optrodes using composites of biological glasses, photo-crosslinkable polymers and genetically encoded molecular sensors. These sensor arrays are designed to measure gradients of extracellular signals (e.g. neurotransmitters, calcium, inflammatory cytokines) and metabolites (e.g. oxygen, glucose, lactate) from tumor spheroids, developing embryos, pancreatic islets and networks of neural cells.
Publications by Year
Visit our Publication Pages at Google Scholar Citations and Research Gate.
- 1. Rickus, J.L., Chang, P., Tobin, A.J., Zink, J.I., Dunn, B. 2004. “Photochemical Coenzyme Regeneration in an Enzymatically Active Optical Material.” Journal of Physical Chemistry B. 108(26) 9325-9332.
- 2. Rickus, J.L. 2005. “Impact of Coenzyme Regeneration on the Performance of an Enzyme Based Optical Biosensor: A Computational Study.” Biosensors and Bioelectronics, 21:965-972.
- 3. Haddish, N.B.; Nyquist, C.; Haghighi, K.; Corvalan, C.; Keshavarzian, A.; Campanella, O.; Rickus, J.; Farhadi, A. 2006. “A multi-scale stochastic drug release model for polymer-coated targeted drug delivery systems” Journal of Controlled Release. 110:314-322.
- 4. Zhao, J., Jedlicka, S.J., Lannu, J.D., Bhunia, A.K., Rickus, J.L. 2006. “Liposome-doped Nanocomposites as Artificial-Cell-Based Biosensors: Detection of Listeriolysin O”, Biotechnology Progress, 22:32-37 (Top 10 Cited Articles in 2006 for Biotechnology Progress)
- 5. Jedlicka, S.S., McKenzie, J.L., Leavesley, S.J., Little, K.M., Webster, T.J., Robinson J.P., Nivens, D.E., Rickus, J.L. 2006. “Sol-Gel Derived Materials as Substrates for Neuronal Differentiation: Effects of Surface Features and Protein Conformation”. Journal of Materials Chemistry. 16:3221-3230 (Featured in Chemical Biology).
- 6. Rickus, J.L. 2006. “Building an Integrated Undergraduate Biological Engineering Program in an Agricultural and Biological Engineering Department: Incorporating the Student Perspective”. International Journal of Engineering Education, 22(1) 53-58.
- 7. P. Banerjee, M.T. Morgan, J.L. Rickus, K. Ragheb, C. Corvalan, J.P. Robinson and A.K. Bhunia 2007. “Survival and growth of hybridoma Ped-2E9 cells under modified culture conditions for potential onsite application in cell-based sensor for Listeria and Bacillus detection” Applied Microbiology and Biotechnology, 73(6): 1423-34.
- 8. S. Padalkar, J. Zhao, K. Stuart, A. Panitch, J. Rickus, L. Stanciu. 2007. “Preparation of Biomolecule Gel Matrices for Electron Microscopy” Ultramicroscopy. 108(4):309-13.
- 9. S.S. Jedlicka, K.M. Little, D.Y. Zemlyanov, D.E. Nivens, J.L. Rickus 2007. “Peptide Ormosils as Cellular Substrates” Journal of Materials Chemistry. 17:5058.
- 10. S.S. Jedlicka, J.L. Rickus, D.Y. Zemlyanov 2007. “Surface Analysis by XPS of Sol-Gels Modified with Covalently Bound Peptides” J. Physical Chemistry. 111(40) 11850-11857
- 11. P. Banerjee, D.Lenz, J. P.Robinson, J.L. Rickus and A.K. Bhunia 2007. “Collagen encapsulated hybridoma cells as biosensors: a simple cell-based sensing system in multi-well plates for rapid detection of Listeria monocytogenes and Bacillus cereus” Laboratory Investigation. 88, 196–206.
- 12. Haddish, N.B., Rickus, J.L., Haghighi, K. 2007. “The role of multiscale computational approaches for rational design of conventional and nanoparticle oral drug delivery systems” International Journal of Nanomedicine, 2(3) 315-331.
- 13. J.J. Koehler, J.Zhao, S.S. Jedlicka, D.M.Porterfield, J.L. Rickus. 2008. “A Compartmentalized Nanocomposite for Dynamic Nitric Oxide Release” J. Physical Chemistry. 1112(47) p. 15086–15093.
- 14. A. Pierce, S. Sommakia, J. Rickus, K. Otto. 2009. “Thin-film sol-gel coatings for neural microelectrodes” J. Neuroscience Methods. 180:106-110.
- 15. S.S. Jedlicka, M.Dadarlat, Y. Lin, A. Young , M. Zhang, T.J Hassell, , P.P. Irazoqui, J.L. Rickus , 2009. “Calibration of Neurotransmitter Release from Neural Cells for Therapeutic Implants.” International Journal of Neural Systems. 19:197-112.
- 16. D Jaroch, M Ward, E Chow, J Rickus, P Irazoqui. 2009. “Magnetic insertion system for flexible electrode implantation.” Journal of Neuroscience Methods. 180:106-110.
- 17. Stratton, J. Rickus, J. Youngblood . 2009. “In vitro biocompatibility studies of the antibacterial copolymer poly(4-vinylpyridine-co-poly(ethylene glycol) methacrylate)-hexylbromide” Biomacromolecules. 10(9):2550-2555.
- 18. Editors: Schachter SC, Guttag J, Schiff SJ, Schomer DL Contributing Authors: Akhtari M, Bailey EM, Baptiste SL, Barkley GL, Beggs JM, Béland R, Besio W, Binder DK, Bonato P, Bromfield E, Brunner P, Carlson C, Cash SS, Cole AJ, Commowick O, Devinsky O, Doyle WK, Dunseath R, Engel J Jr, Eskandar E, Frei MG, French J, Gale K, Gallagher A, Graves N, Gross RE, Guttag J, Halgren E, Hochberg L, Hsu D, Hsu M, Iasemidis LD, Irazoqui P, Koka K, Kuzniecky R, Lassonde M, Lepore F, Ludvig N, Madsen J, Mancinelli C, McGlone F, Medveczky G, Medvedev AV, Morrell M, Nguyen K, Osorio I, Pang T, Patel S, Patritti B, Potter SM, Rickus J, Ritaccio AL, Rogawski MA, Rolston JD, Rotenberg A, Rothman SM, Sackellares JC, Schachter SC, Schalk G, Schiff SJ, Shoeb AH, Schomer DL, Strangman GE, Tang HM, Terry RS Jr, Thesen T, Truccolo W, Ulbert I, Warfield S, Zhang Q. 2009 “Advances in the application of technology to epilepsy: the CIMIT/NIO Epilepsy Innovation Summit.” Epilepsy & Behavior. Sep;16(1):3-46.
- 19. S.S. Jedlicka, J.L. Rickus, D. Zemlyanov. 2010. “Controllable Surface Expression of Bioactive Peptides Incorporated into a Silica Thin Film Matrix” Journal of Physical Chemistry. 114(1). p 342 – 344.
- 20. E.S. McLamore, S. Mohanty, J. Shin, J. Claussen, S.S. Jedlicka, J.L. Rickus, D.M. Porterfield, 2010. “A self-referencing glutamate biosensor for measuring real time neuronal glutamate flux.” Journal of Neuroscience Methods. 189:14–22
- 21. Rajdev, P., M.P. Ward, J. Rickus, R. Worth, and P.P. Irazoqui, Real-time seizure prediction from local field potentials using an adaptive Wiener algorithm. Comput Biol Med, 2010. 40(1): p. 97-108.
- 22. Shi, J., E. McLamore, D. Jaroch, J. Claussen, J. Rickus, and D.M. Porterfield, Oscillatory glucose flux in INS1 pancreatic beta cells: A self-referencing microbiosensor study. Analytical Biochemistry, 2010.
- 23. McLamore, E.S., J. Shi, D. Jaroch, J.C. Claussen, A. Uchida, Y. Jiang, W. Zhang, S.S. Donkin, M.K. Banks, K.K. Buhman, D. Teegarden, J.L. Rickus, and D.M.Porterfield, A self referencing platinum nanoparticle decorated enzyme-based microbiosensor for real time measurement of physiological glucose transport. Biosensors Bioelectronics, 2011. 26(5): p. 2237-45.
- 24. T.G.Cha, B.A. Baker, M.D. Sauffer, Janette Salgado, D.Jaroch, J.L. Rickus, D. Marshall Porterfield, and Jong Hyun Choi. 2011. Optical Nanosensor Architecture for Cell-Signaling Molecules Using DNA Aptamer-Coated Carbon Nanotubes. ACS Nano. 5 (5), pp 4236–4244.
- 25. D.Jaroch, E..McLamore, W.Zhang, J. Shi, J. Garland, M. K. Banks, D. M. Porterfield, J.L.Rickus. 2011. Cell-Mediated Deposition of Porous Silica on Bacterial Biofilms. Biotechnology and Bioengineering. Vol. 108. Issue 10. 2249-2260. Editor’s Choice Selection.
- 26. Shi, J,; Claussen, JC; McLamore, ES; ul Haque, A; Jaroch, D; Diggs, AR; Calvo-Marzal, P; Rickus, JL; Porterfield, DM. 2011. A comparative study of enzyme immobilization strategies for multi-walled carbon nanotube glucose biosensors. Nanotechnology. Vol. 22, Issue 35.
- 27. S.A. Tersey, Y. Nishiki; A. T. Templin; S. M. Cabrera; N.D. Stull, S.C. Colvin, C.Evans-Molina, J.L. Rickus, B.Maier, R.G. Mirmira. 2012. Islet β Cell Endoplasmic Reticulum Stress Precedes the Onset of Type 1 Diabetes in the Non-Obese Diabetic Mouse Model. Diabetes. April 2012 61:818-827.
- 28. Rajtarun Madangopal, Matthew Stensberg, Marshall D. Porterfield, Jenna L. Rickus. 2012. “Directed enzyme deposition via electroactive polymer-based nanomaterials for multi-analyte amperometric biosensors”. Proceedings of the IEEE Sensors 2012. Taipai, Taiwan
- 29. David B Jaroch, Jing Lu, Rajtarun Madangopal, Natalie D Stull, Matthew Stensberg, Jin Shi, Jennifer L Kahn, Ruth Herrera-Perez, Michael Zeitchek, Jennifer Sturgis, J Paul Robinson, Mervin C Yoder, D Marshall Porterfield, Raghavendra G Mirmira, Jenna L Rickus. 2013. “Mouse and Human Islets Survive and Function After Coating by Biosilicification.” American Journal of Physiology. Endocrinology and Metabolism. vol 305, issue 10, pgs E1230-E1240 (PMID:24002572)
- 30. Stensberg MC, Madangopal R, Yale G, Wei Q, Ochoa-Acuna H, Wei A, McLamore ES, Rickus JL, Porterfield DM, Sepulveda M. 2014. “Silver nanoparticle-specific mitotoxicity in Daphnia magna.” Nanotoxicology. Vol. 8, No. 8 , Pages 833-842. PMID: 23927462.
- 31. S.Sommakia, J.Gaire, J.L.Rickus, K.J. Otto. 2014. “Resistive and reactive changes to the impedance of intracortical microelectrodes can be mitigated with polyethylene glycol under acute in vitro and in vivo settings” Frontiers in Neuroengineering. Vol. 7, Article 41, Pages 1-11. (PMCID: PMC4120760)
- 32. S.Sommakia, J.L.Rickus, K.J. Otto. 2014. “Glial cells, but not neurons, exhibit a controllable response to a localized inflammatory microenvironment in vitro.” Frontiers in Neuroengineering. Vol. 7, Article 33, Pages 1-8. PMID:25452724
- 33. Ha, Soo Jung; Gordon Showalter; Shanbao Cai; Haiyan Wang; Wei Michael Liu; Aaron A. Cohen-Gadol; Jann N. Sarkaria; Jenna Rickus; John Springer; Jiri Adamec; Karen E. Pollok; Kari L. Clase. 2014. “Lipidomic Analysis of Glioblastoma Multiforme Using Mass Spectrometry” Current Metabolomics, Volume 2, Number 2, April 2014, pp. 132-143(12)
- 34. Herrera-Perez M, Voytik-Harbin SL, Rickus JL. "Extracellular matrix properties regulate the migratory response of glioblastoma stem cells in 3D culture," Tissue Engineering, Part A. 2015 Oct;21(19-20):2572-82. (PubMed)
- 35. Jennifer L. Kahna, Necla Mine Erena, Osvaldo Campanellaa, Sherry L. Voytik-Harbin and Jenna L. Rickus. “Collagen-fibril matrix properties modulate the kinetics of silica polycondensation to template and direct biomineralization” Journal of Materials Research / Volume 31 / Issue 03 / 2016, pp 311-320. Special Issue on Biomineralization and Biomimetics. (Link to JMR)
- 36. Joerg Rothe, Olivier Frey, Rajtarun Madangopal, Jenna Rickus, Andreas Hierlemann. 2016. “Robust functionalization of large microelectrode arrays by using pulsed potentiostatic deposition”. Sensors 17 (1), 22
- 37. J. Park, M. L. Salmi, W. W. A. Wan Salim, A. Rademacher, B. Wickizer, A. Schooley, J. Benton, A. Cantero, P. F. Argote, M. Ren, M. Zhang, D. M. Porterfield, A. J. Ricco, S. J. Roux and J. L. Rickus. 2017. “An autonomous lab on a chip for space flight calibration of gravity-induced transcellular calcium polarization in single-cell fern spores” Lab on a Chip, 17:1095-1103.
- 38. Kahyaoglu LN, Madangopal R, Park JH, Rickus JL. 2017. “Integration of a Genetically Encoded Calcium Molecular Sensor into Photopolymerizable Hydrogels for Micro-Optrode-Based Sensing” ACS Appl. Mater. Interfaces. 9(37):31557-31567
- 39. Siddarth V. Sridharana, Jose F. Riveraa, James K. Nolanc, Muhammad A. Alama, Jenna L. Rickus, David B. Janes. in press. “On-Chip Microelectrode Array and In-Situ Transient Calibration for Measurement of Transient Concentration Gradients Near Surfaces of 2D Cell Cultures”. Sensors & Actuators: B. Chemical.
- 40. R. Marisol Herrera-Perez, Sherry L. Voytik-Harbin, Jann N. Sarkaria, Karen E. Pollok, Melissa L. Fishel, Jenna L. Rickus. "Presence of stromal cells in a bioengineered tumor microenvironment alters glioblastoma migration and response to STAT3 inhibition”. PLOS ONE. March 22, 2018. https://doi.org/10.1371/journal.pone.0194183